Weekly Recap (March 13, 2026)
Phil Bourne, Anthropic Institute, eLife, AI@UVA / AI in Virginia, NIH genomics tech, AIxBio, AI+Rstudio, legibility, how scientists use Claude Code, how to design antibodies, R updates, new papers.
The University of Virginia School of Data Science lost a legendary scientist this week. UVA Today published a memorial recounting Phil Bourne’s life and impactful career as a scientist and builder. I wrote here earlier this week about how Phil impacted my own career in data science. Phil meant a lot of things to a lot of people, not just those of us here in the School of Data Science.
Ethan Mollick: The Shape of the Thing. Mollick’s latest state-of-play piece argues we've crossed into a new phase where AI agents take on hours of work autonomously, and that recursive self-improvement an explicit item on the roadmap of every major lab. A single chaotic week in late February as a preview of the near future is useful: stock market swings, mass layoffs attributed (dubiously) to AI, and a public Pentagon-Anthropic conflict.
Anthropic launches the Anthropic Institute. Anthropic is consolidating its Frontier Red Team, Societal Impacts, and Economic Research groups under a new umbrella called the Anthropic Institute (anthropic.com/institute), led by co-founder Jack Clark. The announcement reads like a company that believes the next two years will be decisive and wants a dedicated apparatus for telling the world what it’s learning. Anthropic Institute has an open position for an analyst ($295,000 - $345,000 annually).
Niko McCarty: eLife Fallout. An account of the internal tensions that led to Michael Eisen’s firing as EIC of eLife, and what happened to the journal’s radical publishing reforms afterward.
Armin Ronacher: AI And The Ship of Theseus. Interesting line of thought on what AI coding agents mean for re-implementing software for the purposes of re-licensing (e.g. GPL to MIT). See also the discussion on HN.
Ryan Wright and Varun Korisapati: State of AI in the Commonwealth - Global & VA Economic Impact. A new report from the UVA McIntire’s Center for Management IT surveying AI's economic footprint in Virginia, from data center dominance in Northern Virginia to workforce displacement risk and energy grid strain. The policy recommendations are fairly standard, but with a Virginia-specific framing. Read the full report here.
Måns Thulin: Why I don’t use tidymodels. Strong critique of tidymodels: its bootstrap p-values and confidence intervals can contradict each other, and simulations show its bootstrap test actually performs worse than a standard t-test under non-normality. Also missing LOOCV and residual resampling.
Brookings: The train has left the station: Agentic AI and the future of social science research.
New NIH Highlighted Topic: Technology Development for Genomics.
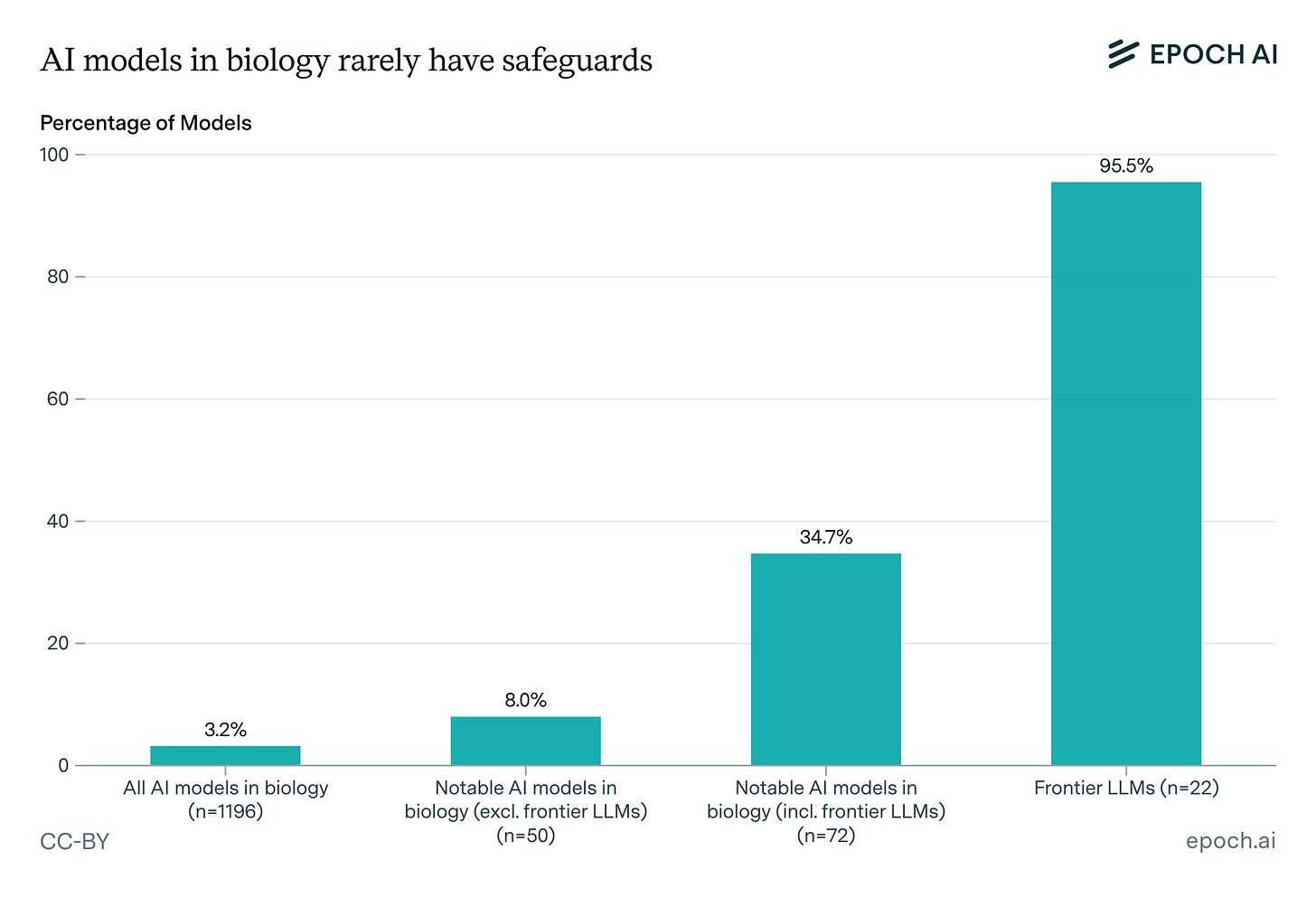
Expanding our analysis of biological AI models. Epoch AI’s expanded database of 1,100+ biological AI models reveals a stark safeguards gap: only 3.2% have any documented safeguards, and just 2.5% underwent pre-release risk assessments, with frontier LLMs accounting for the majority of both. The field remains overwhelmingly academic, dominated by protein engineering and small molecule design, with most models building on a handful of foundation models (ESM-2 chief among them) and progress appearing more data-constrained than compute-constrained.
Joe Cheng, Nick Rohrbaugh, and Sara Altman (Posit): Introducing AI in RStudio. Posit is embedding an AI agent (Posit Assistant) and inline code completions (Next Edit Suggestions) directly into RStudio, backed by a zero-data-retention agreement with Anthropic, at $20/month. The framing is notably honest about their initial skepticism and why they changed their minds. Worth reading if you’re trying to think through when AI assistance in data analysis is and isn't appropriate.
Isabella Velásquez: posit::glimpse() Newsletter – March 2026. This one’s packed full of goodies. Posit AI, built-in PDF viewer in Positron, mirai, Quarto 1.9, Pointblank for Python, learning & community, Claude Code at next week’s Data Science Lab.
Matthew Carter: The Legibility Problem. A thoughtful piece on what happens when AI systems produce scientific discoveries that humans can’t understand or interpret, framed around the chess analogy (we trust engines even when their moves make no sense to us), and what it would take to build the translation infrastructure between AI-generated knowledge and human science.
Julia Silge: Outgrowing your laptop with R and Positron. More details at slides available on the first post on the new Positron blog.
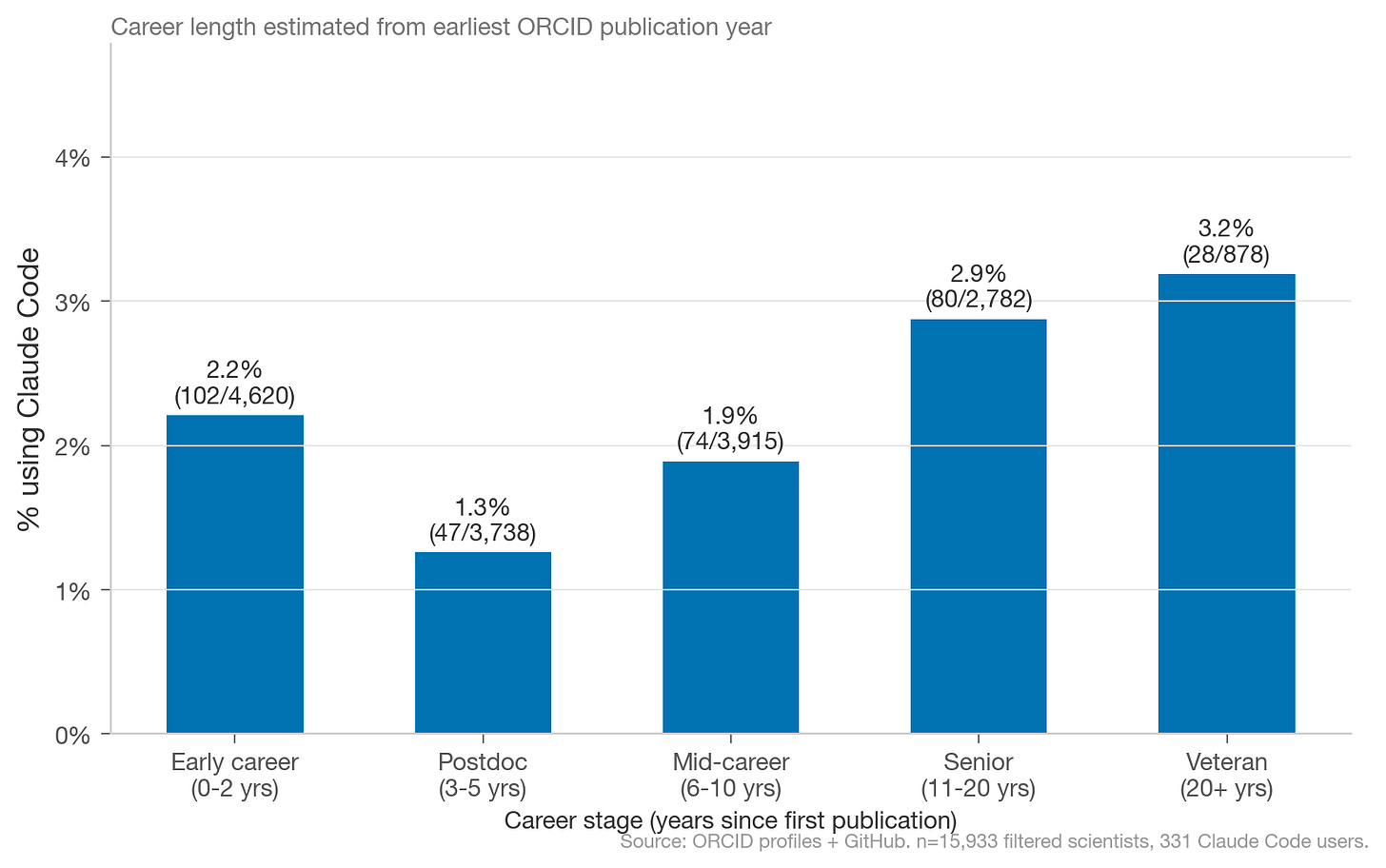
Charles Yang: How Do Scientists Use Claude Code? A clever use of Claude Code’s default co-authorship commits and ORCID-GitHub profile links to measure AI coding tool adoption among ~16,000 computationally active scientists: about 2.1% as of February 2026, with a U-shaped adoption curve by career stage and surprisingly uniform rates across scientific fields. The sample size is small enough that the breakdowns are directional rather than definitive, but this is the kind of empirical work the AI-for-science discussion has been missing. Code for this analysis is here on GitHub.
Matt Lubin: Five Things: March 8, 2026. America vs. Anthropic, Alibaba paper, Ginkgo splits up, biosecurity screening paper, GPT-5.4. I was going to do a short blog post on that biosecurity screening paper, but Matt did so, so I’m not going to. From Matt’s post:
In October, Science published an article led by Bruce Wittmann from Microsoft (and several DNA synthesis companies) demonstrating that AI can design dangerous sequences that evade those companies’ security screening protocols. This past week he and his team followed-up on this work with a new preprint: “The Limits of Sequence-Based Biosecurity Screening Tools in the Age of AI-Assisted Protein Design.” The setup: take dangerous proteins (officially called “proteins of concern”) and use AI to generate “reformulated synthetic homologs”: sequences that do the same thing as the dangerous protein but look different enough to potentially evade screening. This time, they fragmented the sequences into smaller segments (the way someone might actually order them from a synthesis company to avoid detection) and run them through four major biosecurity screening software (BSS) tools.
The results are a mixed bag. Two of the four tools were capable of “robustly detecting fragments as short as 50 nucleotides” — actually exceeding the requirements in the U.S. Framework for Nucleic Acid Synthesis Screening. The other two improved with upgrades. All in all, this is very good news! Of course AI technology is improving constantly, and at some point it might be able to design novel proteins from scratch. Once that occurs, we will need to step up the defensive AI tech to use the same methods to screen novel compounds as one might do to design them. Hopefully someone is working on that.
Devansh: The Real Cost of Running AI. A thorough walkthrough of inference economics, from the FLOPs-per-attention-head level all the way up to the crossover math on edge vs. cloud deployment. Decode is memory-bound, not compute-bound, and that structural mismatch only gets worse with each GPU generation. Long, but helps to understand why output tokens cost more than input tokens, why the KV cache is the real scaling bottleneck, and why every novel architecture is ultimately trying to kill the same quadratic term.
Sayash Kapoor and Arvind Narayanan: Towards a Science of AI Agent Reliability. Borrowing frameworks from aviation and nuclear safety, this post decomposes agent reliability into 12 metrics across 4 dimensions (consistency, robustness, calibration, safety). ~2 years of rapid capability gains have produced only modest reliability improvements. Agents that can solve a task often fail on repeated attempts under identical conditions, and their self-reported confidence is barely better than chance. A good frame for understanding why benchmark progress hasn't translated into economic impact as fast as the vibes would suggest.
R Weekly 2026-W11: Claude skills for R users, beeswarm charts in R, AI in RStudio.
Brian Naughton: How to Design Antibodies: A step-by-step guide to making de novo binders.
Tool Commercial Full Ab VHH Mini-binder Peptide
-----------------------------------------------------------------
BoltzGen ✅ ✅ ✅ ✅ ✅
Germinal ❌ ✅ ✅ ❌ ❌
mBER ✅ ❌ ✅ ❌ ❌
RFantibody ✅ ✅ ✅ ❌ ❌
Mosaic ✅ ✅ ✅ ✅ ✅
BindCraft* ❌ ❌ ❌ ✅ ✅Claus Wilke: PLMs are mostly just memorizing stuff.
New papers & preprints:
Rubrics, Not Vibes: Structured AI Integration as Quality Control for Peer Review
snputils: A High-Performance Python Library for Genetic Variation and Population Structure
geneSTRUCTURE: A Modern Platform for Visualization of Gene Structures
The Limits of Sequence-Based Biosecurity Screening Tools in the Age of AI-Assisted Protein Design
Artificial intelligence-enabled multi-scale virtual cell: perspective, challenges, and opportunities
Gene regulatory networks: from correlative models to causal explanations
Bringing the genetically minimal cell to life on a computer in 4D: Cell